Configuring a Segmentation Mapping Resource for your Model
The labeling of different types of regions within a volume is typically done with "mask" images, images whose pixel values are integers uniquely assigned to the different regions of interest. It is essential to know what the mapping is between region type and mask-image pixel value. There are two components to this mapping: the labels given to region types and the integer value assigned to the region types used in the mask images. You or a colleague will use the OHIF viewer to draw the outlines of organs. During the segmentation process, you will manually enter a label for each organ that you identify (Liver, Spleen, ...). You should identify the set of labels you will use in advance in order to use them consistently in the manual segmentation process. Further, there is a later task that will convert the labeled objects to a mask file where voxels are identified numerically (Liver = 10 Spleen = 20, ...). This mapping should be defined and associated with the model using the following procedure. Note that the value for background is always zero and should not be specified.
1. Create The Mapping File. Create a text file with multiple lines, each containing a region label and its assigned value. For example,
Example mask-value map file contents
Liver=10Spleen=20Kidney=30
Save this file with name "region2MaskValueMap.txt"
2. Attach this file as a model resource in XNAT.
- Navigate to the Project summary page, select the "Machine Learning" tab, and click "Open Machine Learning Dashboard".
- Find your model under "Installed Models" and click on "View Model".
- In the "Actions" box on the Model report page, click on "Manage Files". This will launch a "Manage Files" dialog box.
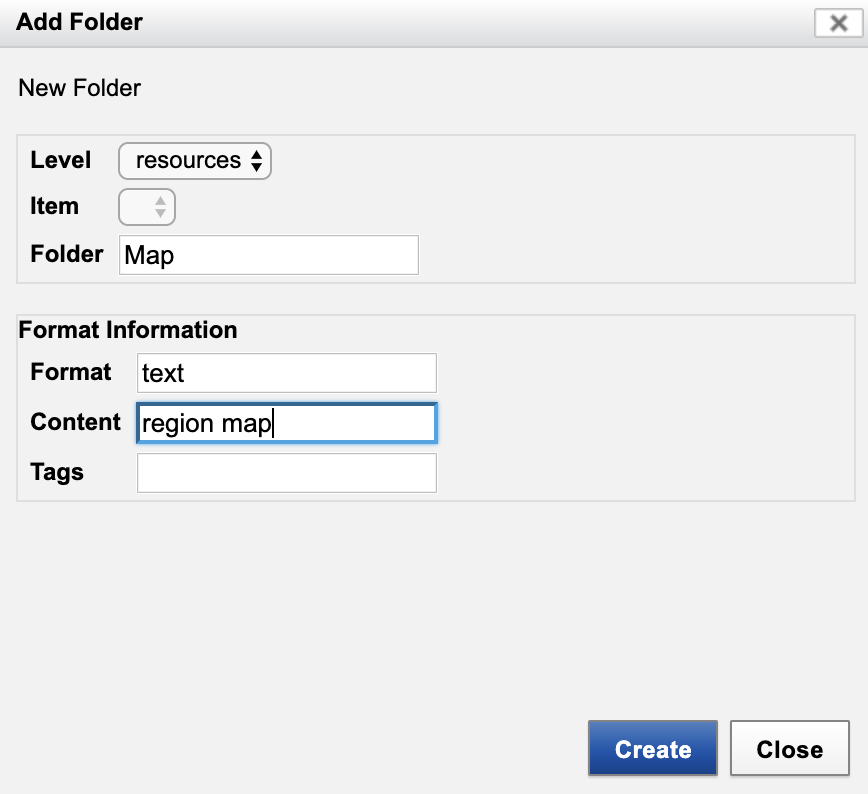
- Click "Add Folder" to launch "Add Folder" dialog.
- Enter as shown. Click "Create".
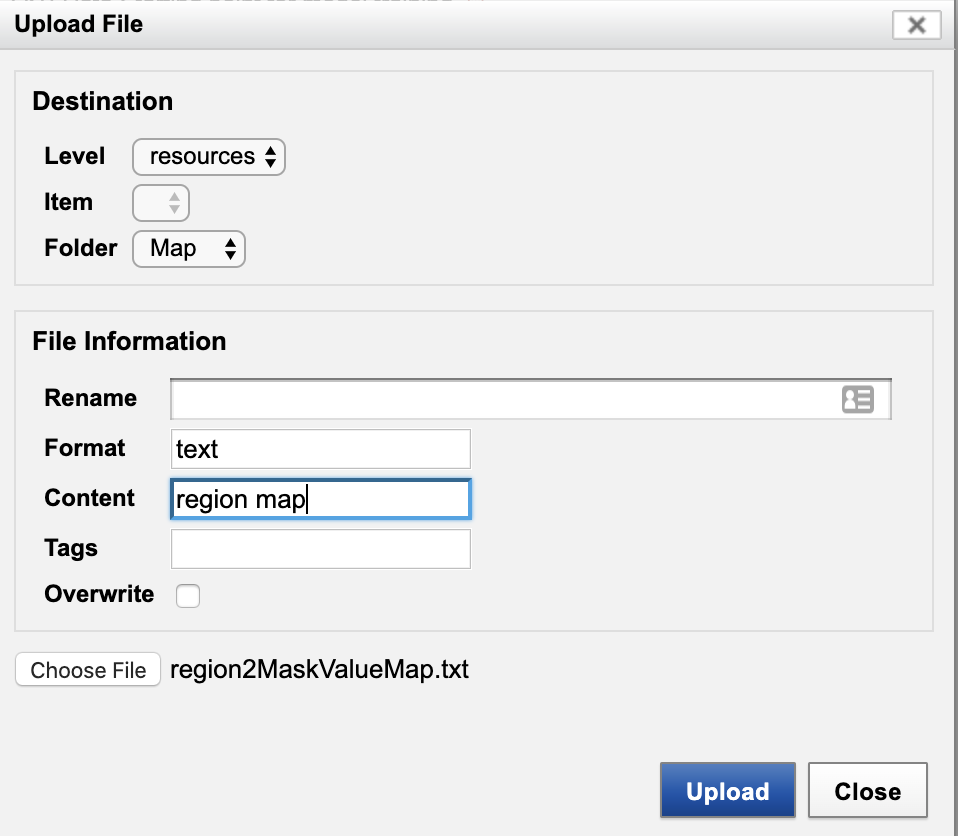
- Back in the "File Manager" dialog, click "Upload Files".
- Enter as shown. Click "Upload".
The region labels and mapping to mask pixel values are now associated with the machine-learning model.