Cancer Imaging Centre, Institute of Cancer Research, UK
OK, guys, so it's more than the few sentences Dan asked for. Hope you don't mind the self-publicity, but I thought it would be useful to let people know in a bit more detail what we do.
About our group:
Together with the Royal Marsden Hospital (RMH), the Institute of Cancer Research (ICR) forms the largest comprehensive cancer centre in Europe, ranked in the top four worldwide.
The CRUK-EPSRC Cancer Imaging Centre is a multidisciplinary group of around 50 staff and students, funded in combination by Britain's leading cancer charity, Cancer Research UK (CRUK) and the Engineering and Physical Sciences Research Council (EPSRC), which is the UK government's primary route for funding research in the physical sciences. The research undertaken ranges from biochemistry, through physics and engineering, to radiology and oncology, working together with approximately a further 50 internal collaborators in closely allied groups from both the ICR and RMH. The Cancer Imaging Centre has developed instrumentation and methodology for MRI and MRS measurements, translating these developments through to clinical applications. To take advantage of these, a wide range of software tools have been created for pre-clinical and clinical use. These software tools are continuing to develop in response to new techniques, and are now being extended to encompass CT, PET/CT and histological comparisons. The group has led reports on standardising DCE-MRI analysis, and is contributing significantly to the standardisation of diffusion MRI analysis in cancer.
 | I'm Simon Doran, the Informatics Lead for the Cancer Imaging Centre. (This was me when I was younger and had a bit more hair!)
|
|---|
My role at the Cancer Imaging Centre involves both definition of the informatics strategy active scientific research. My main interests are:
- development of a "Research PACS" system suitable for use by imaging physicists, biologists and research radiologists;
- translation of research developments into the clinical workflow;
- multimodality imaging (currently MR, CT, PET, SPECT);
- storage and processing of large datasets (e.g., histology, x-ray micro-CT, optical CT: typical data sizes tens of GB per 3-D image);
- image analysis and machine learning;
- quantitative imaging.
My other active research involves technique development for 3-D optical computed tomography. The main application area for this currently is phantom imaging for radiotherapy treatment planning. I am currently very interested in a 3-D microimaging for novel synchrotron-based radiotherapy treatments.
| This video shows a maximum intensity projection (MIP) of a high-resolution 3-D optical computed tomography (CT) image acquired at the University of Surrey, Guildford, UK. The sample is a 9.7 mm-diameter cylinder of a special polyurethane radiation detector known as PRESAGE(TM). The plastic is radiochromic (i.e., changes colour where radiation hits it) and acts like "3-D film". The sample was irradiated at beamline ID17 of the European Synchrotron Radiation Facility, mimicking an advanced cancer treatment technique known as microbeam radiation therapy. Each individual track seen corresponds to an ultra-thin beam of radiation of diameter approximately 50 microns. Optical CT is the only technique that has so far been demonstrated to be capable of making this type of measurement on a 3-D dose pattern. |
|---|
We don't have a formal informatics team at the Cancer Imaging Centre. Rather, our development work is split between me, our system administrator James d'Arcy and a number of post-docs and PhD students.
About our XNAT project(s):
We have a more-or-less standard production system of XNAT 1.5.4 installed, containing (at the time of writing) 16 projects and 355 imaging sessions. The majority of these correspond to Phase 1 clinical trials. However, this is very much the tip of the iceberg, as we have only recently started recruiting users. We are investigating the possibility of translating some of this informatics research into the clinical workflow, which will multiply the data stored by a large factor.
The novel work that we are currently doing with XNAT is in a number of different areas and will be described in more detail in a forthcoming publication in the journal Radiographics (currently scheduled for November 2012):
- incorporation of XNAT into workstation applications;
- extending the range of file types that can be uploaded to XNAT;
- definition of region-of-interest objects to allow seamless transfer of data from diagnosis to therapy;
- XNAT for preclinical data and genomic data;
- visualisation within XNAT.
 |
|---|
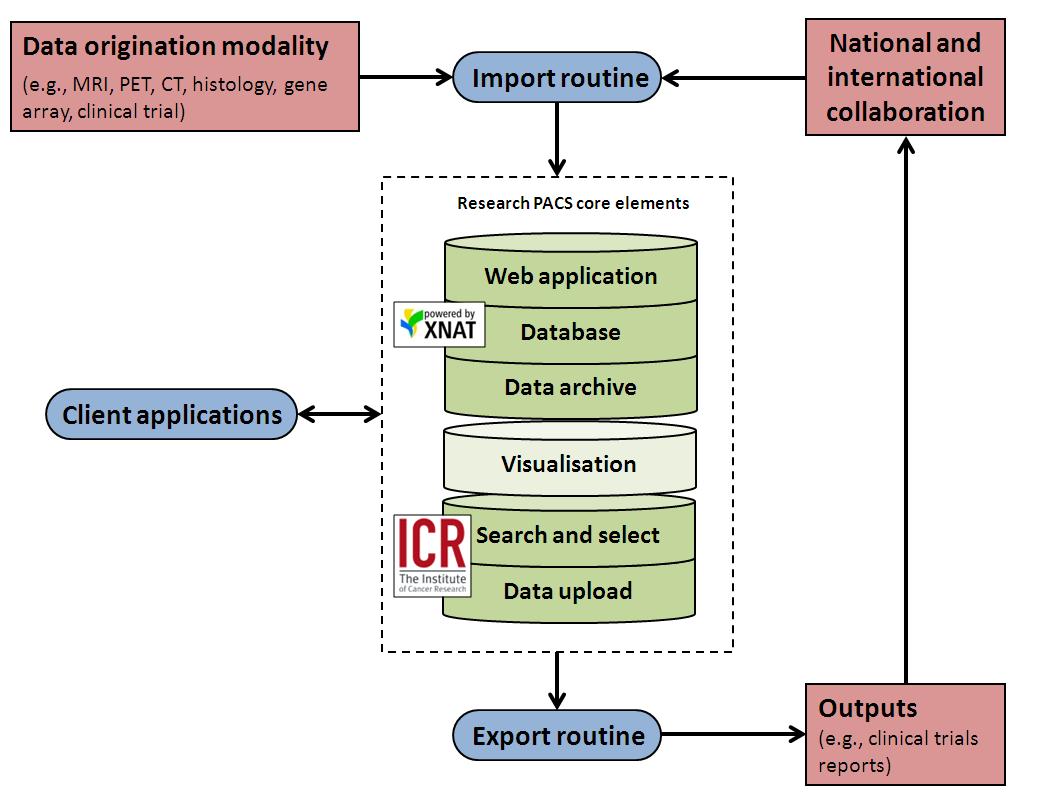
| Figure 1: General schematic of the Cancer Imaging Centre's research PACS, based on XNAT. The visualisation element is shown in a lighter colour as, in our view, the built-in support is not addressed properly within XNAT (at least as of 1.5.4). We are very interested in the potential solutions to fill this gap. |
(1) The ICR DataChooser
We have a legacy of significant numbers of workstation-based applications and a user-base that spends a lot of time developing proof-of-principle code primarily in environments such as IDL (http://www.exelisvis.com) and MATLAB (http://www.mathworks.co.uk/products/matlab). Because of this our use of XNAT perhaps differs from the norm. We see the XNAT webapp primarily as useful for gaining an overview of the data archive, but will interact with XNAT via the REST interface when it comes to downloading the files themselves. The web-browser approach fits well with the general trajectory of radiology applications, namely a gradual move away from traditional fixed PACS workstations towards web-based access to image data, even to the extent of accessing images via mobile devices. It is also similar to the approach adopted by online image archives such as those maintained by the National Biomedical Imaging Archive (NBIA), which allow users to both browse and perform sophisticated searches. However, the end product of these interactions is generally a download to a potentially non-managed file path on the user’s own workstation. This concerns us because of the potential for data duplication and/or for data modification or corruption (often unintentional and sometimes unnoticed — users think they are looking at the original data, when, in fact, some application on the local workstation has changed it). Given that the motivation for image downloads is generally to analyse data using some locally installed piece of software, it makes far more sense to allow applications to interact directly with XNAT. Thus, one of the major aims of our new piece of work is to make all data transfer to the local machine seamless, storing files (where necessary) in a well-structured cache which is, as far as possible, invisible to the end user.
 |
|---|
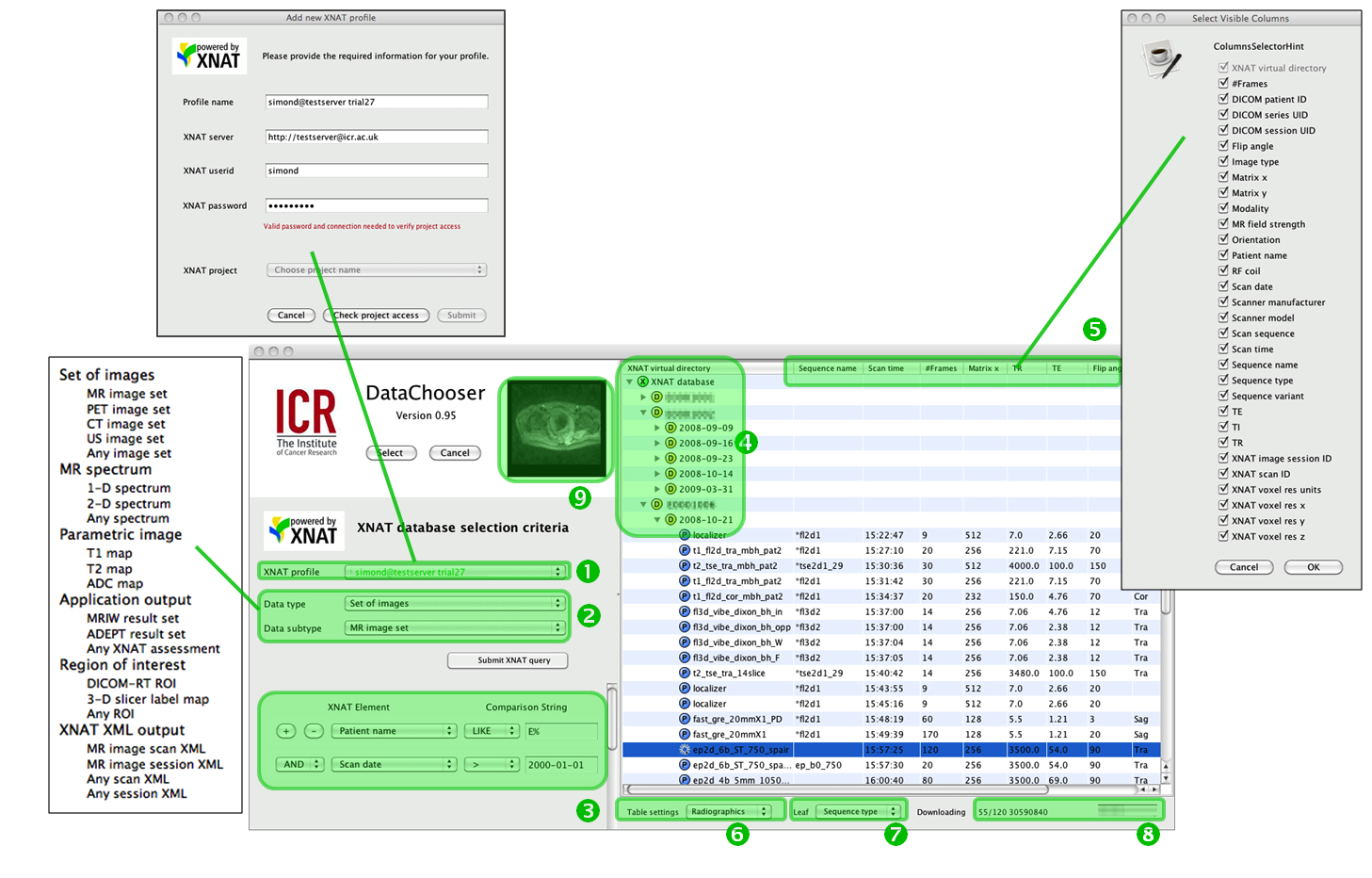
| Figure 2: The DataChooser interface: (1) profile selector and management; (2) return type and subtype selector, including datatypes associated with the local applications Magnetic Resonance Imaging Workbench (MRIW) [18] and Adept Data Exploration and Processing Tool (ADEPT); (3) data selection rules; (4) virtual directory system; (5) current metadata selections (table header line) and popup allowing changes; (6) table settings manager; (7) leaf-element selector (see main text); (8) download progress; (9) cine image preview. Note: In this and subsequent figures, any identifiable names have been deliberately obscured. |
- The Data Chooser manages multiple simultaneous connections to an arbitrary number of XNAT database servers by means of a set of profiles.
- The Data Chooser maintains a local definition of the high-level data types that this particular installation “knows about” and can return to users. Users choose one of these and the search results displayed will reflect all the items of this type accessible to the selected XNAT profile and matching the search criteria. Types such as DICOM images will be common to all institutions, but the system is aware of local data types, too.
- Results are returned using a simple implementation of the XNAT search mechanism, with easy specification of up to 8 chained conditions.
- Virtual directory listing: Although XNAT does has its inbuilt data hierarchy it is often useful to sort results into arbitrary hierarchies. The system uses the criteria specified in (3) to establish a virtual directory structure, which is displayed as the “tree” section of a “tree-table”. Whilst a typical clinical view of the database might arrange results in the form patient name -> date, as in Figure 2, a study comparing similar protocols on different scanners and coils might, for example, wish to arrange results by manufacturer -> scanner model -> RF coil, as shown in Figure 3. To create this figure, the Data Chooser was “pointed at” the public domain repository http://central.xnat.org and re-ordered the entire publicly viewable contents in around 5s. The icons with coloured letters represent different classes of information: for example, “E” codes for information related to the equipment (in this case scanner manufacturer); “D” codes for demographic information (patient identifier); “P” for experimental parameters; “U” represents a DICOM UID, and so on.
- Each selectable item in the repository is associated with metadata. Whilst not all DICOM tags are carried forward into the XNAT database, many are present and any combination of these can be displayed in the “table part” of the tree-table browser, by dragging columns and choosing the elements to display in the “Select visible columns” popup.
- The table settings specified via (5) above can be stored on a per-user and a per-data type basis.
- We realised that it does not always make sense to use the same “leaf” type for the tree table. For example, it is often highly convenient, as in Figure 2, to make the leaves display the MR pulse sequence description coded into the DICOM files. However, on some occasions, the same sequence is run many times sequentially (for example, in a time-course experiment). Under these circumstances, one sees many rows of the table that look superficially identical. Thus, we allow the user to choose an alternate label for the leaves (e.g., the scan time or the series number) to facilitate selection.
- Download monitor: The system works equally well whether the images are stored on the local machine that is running the Data Chooser itself or on a remote repository (e.g., the XNAT Central server in Figure 4). The only difference is the speed of image retrieval.
- Data preview: When the user clicks on an individual item in the virtual directory listing, a representation of that item is displayed in the preview box. For a set of images (DICOM series), this takes the form of a cine loop, whereas when other data types are being selected, the system will present an appropriately representative preview.
 |
|---|
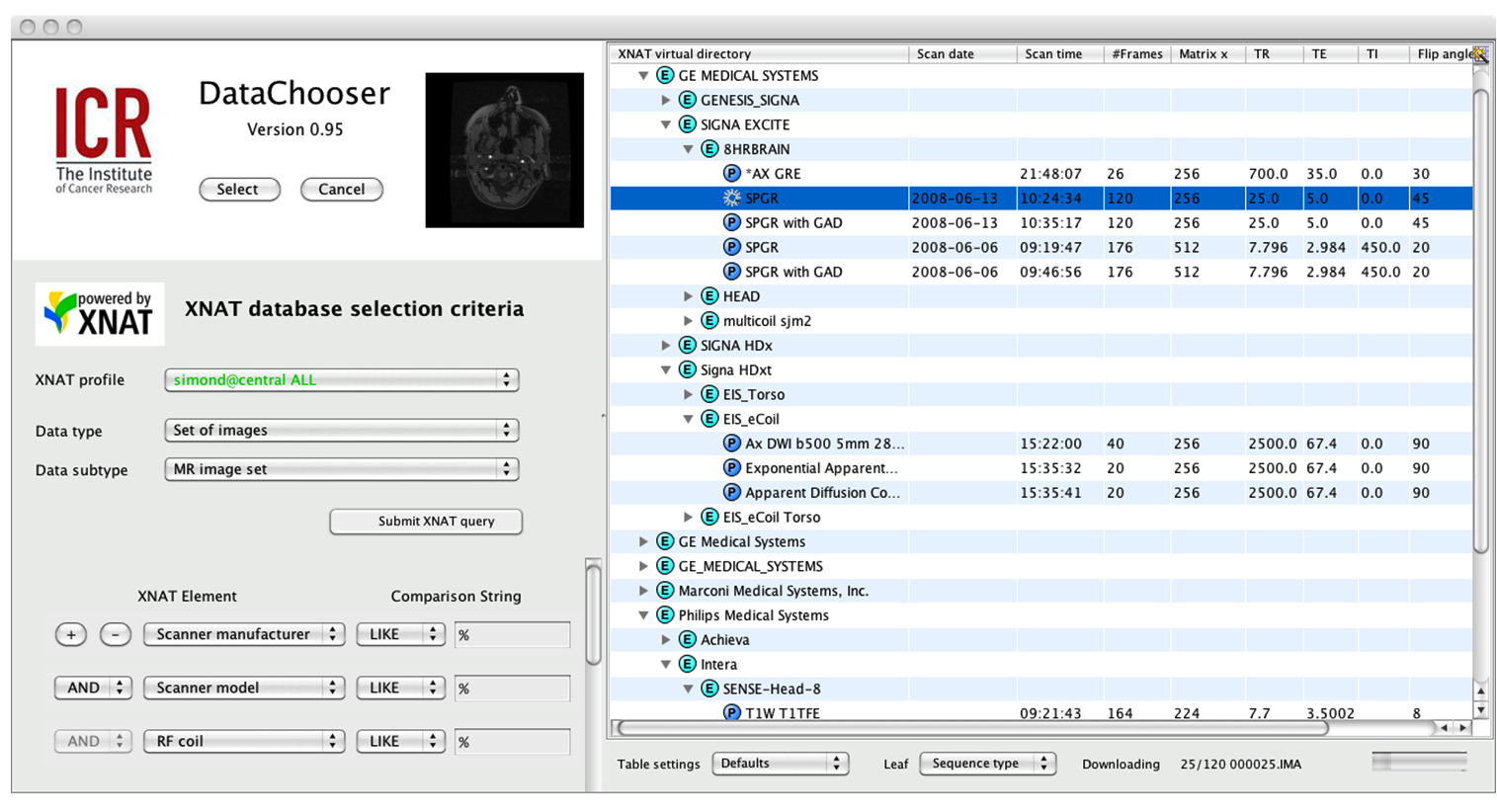
| Figure 3: Data Chooser used to access a public domain XNAT repository (http://central.xnat.org) and demonstrating how an alternate set of selection criteria lead to a different virtual directory structure |
(2) The ICR DataUploader
This is the converse of the Data Chooser, and is equally easy to integrate within an application. It provides a way of inserting files of any of the locally known data types into XNAT. A user selects a data type and a data file from the local (or networked) file system. The uploader parses the file to verify that it conforms in format to what is expected in the XNAT schema and extracts the metadata needed to populate the Postgres database. This is still something of a work in progress: it is very slow compared with the NRG DICOM uploaders and we are still extending the number of types that can be uploaded. However, since we subclass the parser for each datatype that we handle, it does offer significant flexibility in enforcing arbitrary conditions on the uploaded data. For example, for one of the local types, which is represented by XML files that include references to the DICOM files of the images processed, we choose to enforce the condition that the new data file cannot be uploaded to the repository unless the dependency images are already there. We then ensure that the new data are automatically uploaded to the correct XNAT session.
 |
|---|
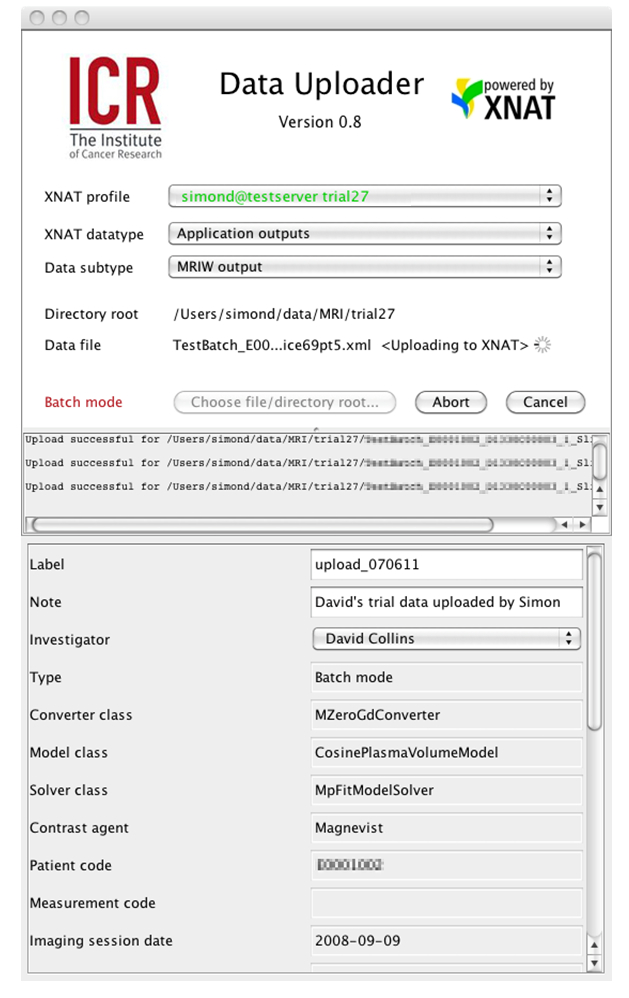
| Figure 4: The ICR Data Uploader interface. Note how metadata elements from the XNAT XML schema corresponding to this DC-MRI datatype created by MRIW are parsed from the uploaded file and displayed to the user. |
(3) Regions-of-Interest
One of our major clinical imperatives is to be able to take images from our scanners, analyse them in proprietary and in-house software, select regions-of-interest (ROIs) defining tumours and then transfer these ROIs to the radiotherapy treatment planning computers. The latter use the DICOM-RT extensions to specify ROIs. This is a very rich and flexible, but somewhat complex, nested format. I have created an XNAT-compatible schema to allow DICOM RTSTRUCT files to be uploaded to XNAT and searched via the ICR DataChooser. Again, this is a work in progress, which I hope to develop over the coming months.
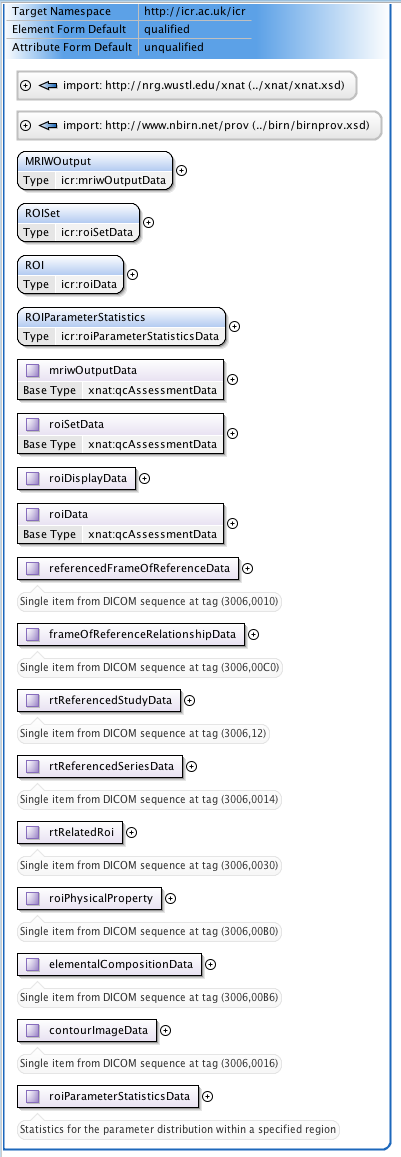 | 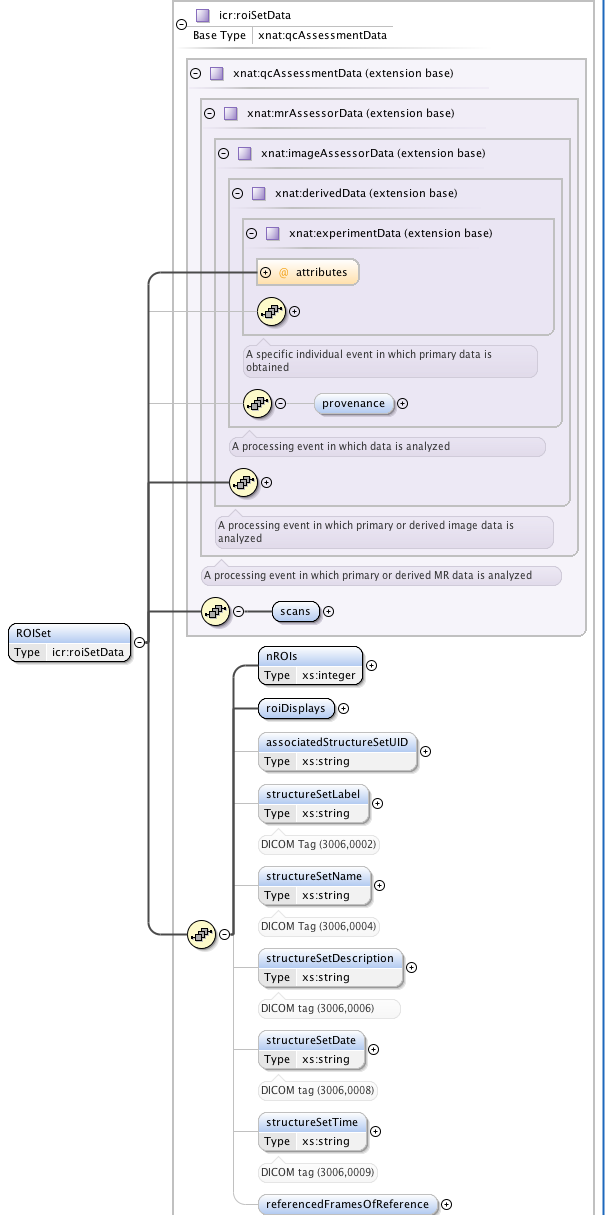 |
|---|---|
| Figure 5: Part of the extensive schematic created to implement | the DICOM RTSTRUCT file definition into XNAT |
(4) Preclinical PACS and radiogenomics
A significant portion of the work that goes on in the unit is preclinical. We wish to have a common research PACS interface for managing both clinical and preclinical data. As part of this, it will be necessary, over the coming months to add extensions to the subject demographics schema element to code for all the properties associated with non-human subjects and I hope to build on the work previously reported by Kasper Claes (https://groups.google.com/forum/?fromgroups#!searchin/xnat_discussion/Kasper/xnat_discussion/0ivHehQHmaY/gx-rKX0w5o4J). We are also keen to add display of histology images to the system and eventually genetic data, too.
(5) Visualisation
Although XNAT is not designed as a PACS replacement per se, it seems to us that with clinical PACS evolving web-based viewing solutions, the time is right to consider bundling XNAT with a much enhanced viewing interface. We are very interested to discuss what might be possible here, as when our radiologist colleagues have encountered commercial software that allows genuine PACS-quality visualisation from a web browser on a high-end monitor, they have been very excited.
Our goals for the XNAT Workshop:
Learn lots and have a great time ...
Specifically:
- I am particularly interested to find out more about customising web report pages to include graphics and visualisations. I was very impressed with Gabriele's post https://groups.google.com/forum/?fromgroups#!searchin/xnat_discussion/Gabriele/xnat_discussion/aN4RC9YUmPo/bL9DVpn8rqgJ and would like to learn how to do this sort of cool stuff!
- I'm also excited by the XNAT Marketplace concept.
- As discussed above, I am keen to talk about visualisation and data viewing. This will make a big difference to take-up by our research radiologists.