Database Management Unit: Max Planck Institute for Human Cognitive and Brain Sciences, Leipzig
About our group:
The Database Management Unit (DBU) at the Max Planck Institute for Human Cognitive and Brain Sciences in Leipzig (MPI CBS) is responsible for the development of scientific databases and other resource-management and discovery tools used by our various scientific groups.
The DBU oversees the day-to-day running and maintenance of a number of active databases of patients, adult and children test subjects, and studies conducted at the institute. We maintain a number of systems; amongst them, a subject management and recruitment system with data about our experimental subjects, and a content management system with critical information about our scientific experiments. The unit has currently a workforce of two main developers, some data QC and data entry staff and me, Roberto Cozatl, as leader/coordinator of the unit. The XNAT development work has been done by Gabriel Rivera.
About our XNAT project(s):
At the MPI CBS, we have deployed an XNAT 1.5 installation that currently contains data for four projects and about 120 MR sessions. Through this pilot project, we aim to evaluate how XNAT can be used to support our established methodologies for data management, processing pipelines, and data sharing procedures. This, not only between different research groups at the institute, but also between our existing collaborators in other institutions. This pilot installation was made taking into account the research focus of two projects in particular:
Project 1: Maxnet Cognition XNAT/ XNAT Sync Manager
One of the areas of research at the Department of Neurology of the MPI CBS - led by Professor Arno Villringer http://www.cbs.mpg.de/depts/n-3 - concerns the assessment of genetic influences on brain plasticity. In this context, the department is a member of the MaxNet Genetics and Cognition initiative which is a collaborative project aiming at the discovery and replication of relations among genes, brain, and behavior via the sharing and analysis of MR and genetics data over multiple centres (currently 3 participating centres). GWAS genomic data of many subjects have been gathered for this project. Now, and after a lot of legwork done to agree on a standard imaging protocol amongst participating centres, MR data sets – consisting mostly of T1s, DWI, and resting state data – are now being added to the project. The amount and rate of data acquisition planed for this project will require a sound management and data sharing approach and state-of-the-art technical solutions.
After some investigation of XNAT’s capabilities, we determined that it would be the perfect tool to facilitate our goals, due to its extensible design and very robust web application, which is ideal for internet data transfer for our multi-centre project. Initially, the deployment of a single centralized XNAT site to support the project was considered, but there were several benefits to having a separate installation at each institute. With a local XNAT, each participant can also use XNAT for non-Maxnet projects, and perform their own user access control and IT administration. Additionally, the benefit of having LAN data transfer speeds while moving data in and out of the system was considered.
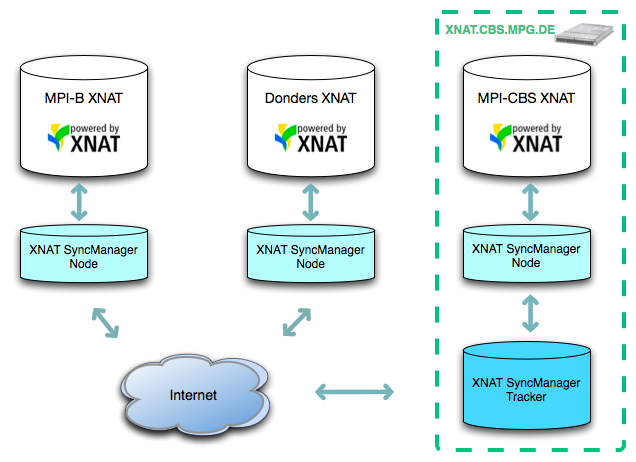
Due to these benefits, the decision was made to roll out an XNAT installation at each centre, and implement a tool to sync XNAT data between the separate servers in the background, through web services that access the REST API through PyXNAT. The result is effectively a small-scale content distribution system for multiple XNAT sites. The sync system, referred to as the Maxnet Cognition XNAT Sync Manager, consists of two components—the Node component that runs on each XNAT Server, and the Tracker component that is currently hosted at the MPI CBS.
The Tracker component serves as a centralized database that stores the REST resource paths, and modification times of the subjects, experiments, and metadata that comprise each shared XNAT project. The Node component, running on each server, performs two main tasks continuously—it iterates through the Maxnet projects to check for new data added locally, and queries the Tracker for any new data that has been added to the shared projects by the other centres. When the Node receives information about new XNAT data from the Tracker, it then uses PyXNAT to download the new data from the server it originated on, and import these data into the local server. Both components run as background daemon processes as a companion to the XNAT installation.
Project 2: Pipelining for Neuroanatomy and Connectivity
In collaboration with Dr Daniel Margulies, Group Leader of the Max Planck Research Group "Neuroanatomy & Connectivity http://www.cbs.mpg.de/groups/misc/nac, we aim to use our XNAT installation in Leipzig as a repository for the sharing of MR data sets that have been created or derived via novel methodologies and or via the use of sophisticated neuroimaging software packages. Our goal is to use XNAT’s features for organizing raw or minimally processed data sets with the corresponding reconstructions outputted from a number of processing pipelines currently being implemented.
One main focus of this work is to integrate pipelines currently built with Matlab-related tools, BASH scripting, and other tools into a format that is compatible with the XNAT Pipeline Engine. We are also working to integrate NiPype python-based pipelines into XNAT through the use of the xnnppx connector. We will continue to focus on NiPype as a comprehensive integration tool for pipelining so interoperability between XNAT and this tool will be beneficial. Other goals here are: (1) the creation of custom schemas in XNAT that describe the provenance of processed data generated by our pipelines, and (2) the sharing of this data between our Maxnet servers using the Maxnet sync manager system described above.
What brings you to the 2012 XNAT Workshop?
We clearly think this is a great opportunity to learn more about XNAT and what others are up to with their development work. We would like to use the opportunity to be in direct communication with XNAT administrators and developers to gain practical knowledge on administration tasks and good inspiration for our next development phase.
I also look forward to hearing more details about the future development of XNAT. Particularly, to what extend the integration of genetic data will be supported in subsequent versions and what might the time line for these developments be.
What do you want to focus on while you're here?
I would personally like to focus on getting first hand advice and knowledge about the most important administration tasks. Unfortunately, due to future work plans and commitments, Gabriel Rivera will leave us soon and will not attend the workshop. Although I will be providing administration support for our XNAT projects, I am a biologist by training, so for some of the more technical issues I will be taking lots of notes to ensure I can pass as much useful information as possible to the person replacing Gabriel in the future. We would like to ensure that our XNAT installation(s) remain in good shape so that further development can take place soon.
In addition to this, I would also like to present the work Gabriel has done so far with our sync-manager to others who might be interested in this topic. I will definitely be interested in learning about other data syncing approaches and discussing data modeling issues with Dave Gutman, Dan Marcus, Tim Olsen and all those signing for the XNAT to XNAT copy-sync sub-project.
...